Where do you start?
The code is freely available and easy to install in a few clicks with Anaconda (and pypi). Please see instructions here. We also provide a very easy to use GUI interface, and a step-by-step user guide!
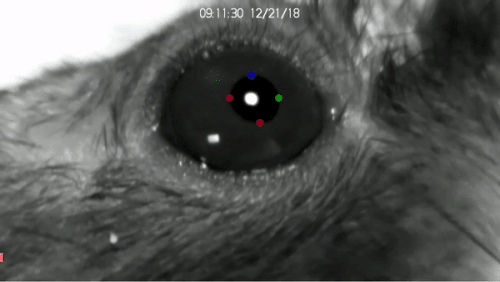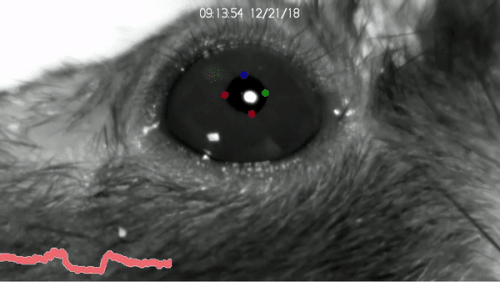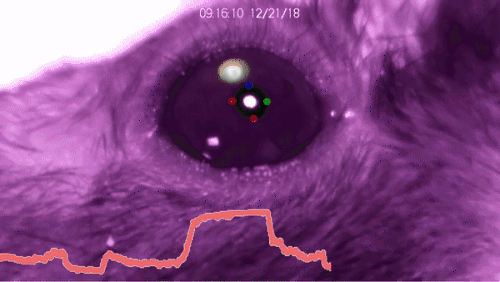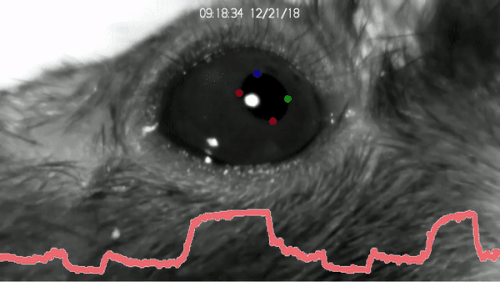
DeepLabCut™ is an efficient method for 2D and 3D markerless pose estimation based on transfer learning with deep neural networks that achieves excellent results (i.e. you can match human labeling accuracy) with minimal training data (typically 50-200 frames). We demonstrate the versatility of this framework by tracking various body parts in multiple species across a broad collection of behaviors. The package is open source, fast, robust, and can be used to compute 3D pose estimates or for multi-animals. Please see the original paper and the latest work below! This package is collaboratively developed by the Mathis Group & Mathis Lab at EPFL (releases prior to 2.1.9 were developed at Harvard University).
⭐️ If you like this project, please consider giving us a star on GitHub! ⭐️
Learn More:
Support DeepLabCut:
In the News:
Key Publications:
Primary research papers:
Mathis et al, Nature Neuroscience 2018 or free link: rdcu.be/4Rep
Nath*, Mathis* et al, Nature Protocols 2019 or free link: https://rdcu.be/bHpHN
Kane et al, eLife 2020
Lauer et al. Nature Methods 2022
Review & educational papers:
Mathis & Mathis Deep learning tools for the measurement of animal behavior in neuroscience Current Opinion in Neurobiology 2020
Mathis, Schneider, Lauer, Mathis A Primer on Motion Capture with Deep Learning: Principles, Pitfalls, and Perspectives Neuron 2020
Preprints/always open access versions:
[1] arXiv (April 2018): Markerless tracking of user-defined features with deep learning (published in Nature Neuroscience)
[2] bioRxiv(Oct 2018):On the inference speed and video-compression robustness of DeepLabCut
(presented at COSYNE 2019)
[3] bioRxiv (Nov 2018): Using DeepLabCut for 3D markerless pose estimation across species and behaviors (published in Nature Protocols)
[4] arXiv (Sept 2019): Pretraining boosts out-of-domain robustness for pose estimation
[5] arXiv (Oct 2019): Deep learning tools for the measurement of animal behavior in neuroscience (published in Current Opinion in Neurobiology)
[6] arXiv (Sept 2020): A Primer on Motion Capture with Deep Learning: Principles, Pitfalls, and Perspectives (Published in Neuron)
[7] arXiv (Nov 2020): Pretraining boosts out-of-domain robustness for pose estimation (published at WACV 2021 & ICML workshop 2020)
[8] bioRxiv (August 2020): Real-time, low-latency closed-loop feedback using markerless posture tracking (published in eLife)
[9] bioRxiv (April 2021) Multi-animal pose estimation and tracking with DeepLabCut (published in Nature Methods)
What’s New:
July 2023! We released an updated preprint and new SuperAnimals into the Model Zoo, see modelzoo.deeplabcut.org for more information!
April 2022! We released a major update to the code with strong multi-animal support, tracking, and many new tools and features! Please check out the new docs and video tutorials to get started, and it’s now published in Nature Methods!
March 2020! We released the DLC Model Zoo. Use trained networks without any installation, coding, or retraining: DeepLabCut Model Zoo homepage
Are you using DeepLabCut?
Please give us feedback (or just a shout out that you are using it) to help us support the project. We appreciate it! Join the 500+ Universities that have added their name!
Get Inspired! Check out who is citing us: